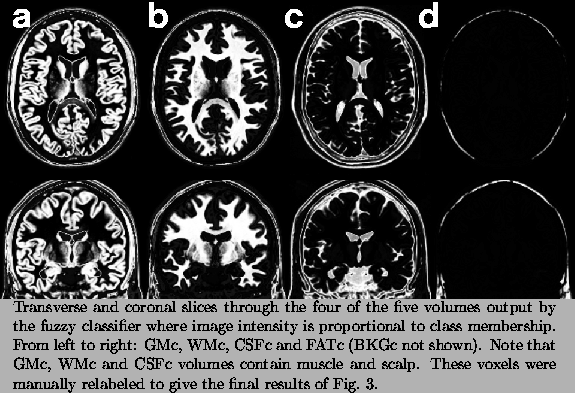
 |
The five volumes (GMc, WMc, CSFc, FATc and BKGc) generated by the classifier (see Fig. 2) contained erroneously labelled voxels and thus required a number of manual and semi-automatic interventions. The result of this corrective process was a 3D tissue model, with one volume per class, where voxel intensity represents the fraction of tissue (between 0.0 and 1.0) within the voxel. Four of the ten volumes that define the brain phantom are shown in Fig. 3. Table I summarizes the correction steps that are described in detail below.
| structure | description | ||
| GM | = | e(GMc |
grey matter within the brain parenchyma |
| WM | = | e(WMc |
white matter within the brain parenchyma |
| CSF | = | e(CSFc |
cerebro spinal fluid surrounding the brain and within the ventricles |
| GL | = | e((GMc
|
layer of glial tissue lining the ventricles |
| M+S | = | e(GMc |
muscle and skin |
| OTH | = | e(WMc |
other tissue |
| FAT | = | e(FATc |
fatty tissue |
| SKN | = | e(CSFc |
mostly skin |
| SKL | = | e((CSFc |
skull (does not include sinuses) |
| AIR | = | e((BKGc
|
air outside head and within sinuses |

As it was necessary to separate brain from non-brain structures, the first
step in the correction phase was the creation of a discrete brain mask (B)
by manual voxel painting using software developed in
house![]() that
permits tri-plane (coronal, sagittal and transverse) roaming through the
volume with arbitrary pan and zoom. The brain mask, which only identified
brain parenchyma and omitted all non-ventricular CSF spaces, was dilated
and further edited by hand to create a second mask (B+) containing all
structures (e.g., brain parenchyma, arteries, dura, sub-dural CSF, CSF-filled
cisterns) within the skull cavity. The complement of each volume,
B
that
permits tri-plane (coronal, sagittal and transverse) roaming through the
volume with arbitrary pan and zoom. The brain mask, which only identified
brain parenchyma and omitted all non-ventricular CSF spaces, was dilated
and further edited by hand to create a second mask (B+) containing all
structures (e.g., brain parenchyma, arteries, dura, sub-dural CSF, CSF-filled
cisterns) within the skull cavity. The complement of each volume,
B![]() and (B+)
and (B+)![]() respectively, were also created to identify
non-brain structures.
respectively, were also created to identify
non-brain structures.
The brain masks were used to separate each of the five volumes into brain and non-brain volumes. In each of following steps, where masking and manual corrections were interleaved, the total proportion of each tissue class in each voxel was kept constant; those removed from one tissue class were re-attributed to other volumes so that the sum of all tissue fractions at each voxel was normalized to 1.0.
The WMc voxels within the brain mask became the final WM phantom volume after manual editing (Fig. 3b), those outside the dilated brain mask (B+) form the other (OTH) class. The remaining WMc voxels (WMc - WM - OTH) were due to partial volume effects between grey matter tissue and CSF. Since these voxels had a maximum value of less than 0.07 (where pure tissue is 1.0), they were ignored and zeroed instead of reattributing them to the GM or CSF volumes.
The GMc voxels within the brain became the final GM class after manual editing (Fig. 3a). The remaining GMc voxels fell into two groups: 1) those within the brain that were manually removed were from the inner surface of the ventricles and were used to form the glial (GL) class; 2) those outside the brain became muscle and skin (M+S).
The fat (FAT) volume was created by editing FATc, using semi-automated region growing and manual painting. The remaining voxels were zeroed. Similar tools were used to produce the the skull (SKL) volume by editing BKGc to identify all voxels within the skull. All BKGc voxels within the brain mask were zeroed. The remaining voxels formed the air (AIR) class.
After manual editing, voxels from the CSFc within B+ formed the final CSF volume containing both ventricular and subarachnoid CSF (Fig. 3c). The CSFc voxels leftover within the brain mask were ignored (zeroed) since they were caused by partial volume regions where grey and white matter tissue occupy the same voxel as well as partial volume regions around the ventricles. (These voxels had maximum CSFc values of 0.02 and 0.03, respectively.) Those that were leftover outside the brain mask were either manually added to the SKL volume or formed the skin (SKN) volume, depending on their proximity to these structures.
Since some voxels from the WMc, CSFc, FATc and BKGc were zeroed, the integral of all tissue components was not equal to 1.0 for all voxels within the phantom. These voxels were simply normalized on a voxel by voxel basis, by dividing their respective tissue components by their sum.
These ten ``fuzzy'' volumes, one for each class, define the digital phantom. Three of those that define the brain are shown in Fig. 3. A discrete version of the phantom was also created by storing the label of the most probable class at each voxel location and is shown in Fig. 3d. A total of 20-30 man-hours were required for all manual intervention required to build the phantom. It is important to note that while there may remain residual errors in the classification of some voxels of the phantom even after the manual correction process, these do not dilute the purity of the phantom for validation studies. By definition, ``truth'' in the tissue labelling for any voxel is what is stored in the reference volumes. To illustrate, a perfect MRI tissue segmentation algorithm should be able to take a simulated MRI image based on the reference volumes and recover the class occupancy of every voxel in the reference volumes.