Georges Le Goualher
Montreal Neurological Institute
McConnell Brain Imaging Centre
3801 University Street
Montreal, Canada H3A 2B4
e-mail : georges@bic.mni.mcgill.ca
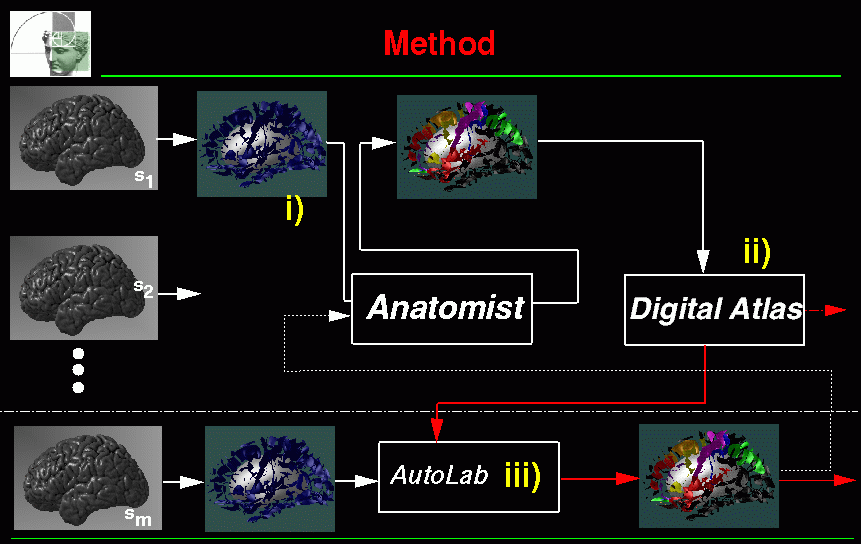
SEAL stands for Sulcus Extraction and Assisted Labelling. Figure 1 shows the overview of the application. There are three main parts :
For data already available and work done within the ICBM project refer to section 6. For related Papers and Reports refer to section 10.

SEAL is a perl script :
~georges/public_bin/sulcus_graph.pl
command:
sulcus_graph.pl final_t1.mnc surface.obj classification.mnc output_dir
3D T1 MRI volume registered and resampled in the MNI-Talairach coordinate system (c.f http://www.bic.mni.mcgill.ca/~louis/mri_segmentation/ , [2]).
Ouput of cortical surface segmentation (i.e. output of ~david/public_bin/cortical_surface.pl, [3] or grey matter surface output of ~georges/public_bin/cortical_surface_double_hull.pl).
Ouput of tissue classication (i.e. output of classify or classify_clean, [5]).
Directory where ouput of sulcus_graph.pl should be stored.
NOTE : sulcus_graph.pl has been applied to the ICBM database.
Output are stored under : /data/icbm/mri1/subjects/00XXX/seg/auto/mni_icbm_00XXX_t1_sulci sulcus_graph.pl generates 6 object files and one sulgraph
file which are stored under the output_dir directory 1.
As an example lets consider ICBM 00100 subject. For this subject, output
files are stored under : The .sulgraph structure is an attributed relational graph structure
that I've defined for my application 2.
A graph contains two main features : Arcs and Vertices 3.
For informations on this ICBM database refer to :
http://www.bic.mni.mcgill.ca/~louis/icbm/icbm_summary.html
Output Data
:
/data/icbm/mri1/subjects/00100/seg/auto/mni_icbm_00100_t1_sulci
These ouput files are :
GRAPH_sulci.sulgraph :
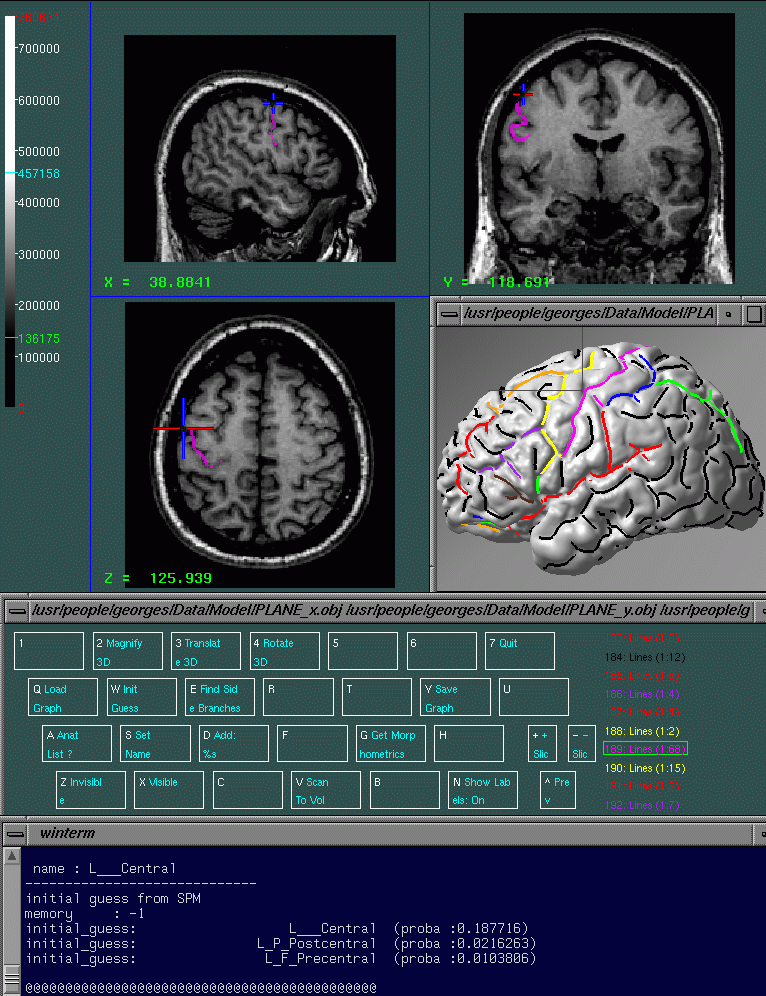
The goal of the Interactive Labelling is to label the sulci of interest. Once this labelling has been performed on several individuals, it's possible to compute sulcus SP_AMs. The interactive labelling is performed using an modified version of Display called Display.geo (see figure 2). The only difference between Display and Display.geo is that Display.geo contains a specific menu for sulcal labelling 4.
You may use directly ~georges/Source/Display/Display.geo for interactive labelling, but I strongly suggest you create a disp.geo alias :
alias disp.geo \verb|~|georges/Source/Display/Display.geo \ ~georges/Data/Model/PLANE_x.obj \ ~georges/Data/Model/PLANE_y.obj \ ~georges/Data/Model/PLANE_z.obj \ ~georges/Data/Model/little_cortex.obj'
Then when you start the command disp.geo you can see that this command start Display.geo and that in this version there is a new button in the main menu : 3D Sulci (associated with the N keyboard key). Several geometrical objects are launched in the 3D window : 3 perpendicular planes and a little cortex model. These objects will be useful to prevent visual overlapping between hemispheres. Of course you can hide an object by :
You need :
Run :
disp.geo final_t1.mnc surface.obj G_A_L_L.obj G_A_R_L.obj
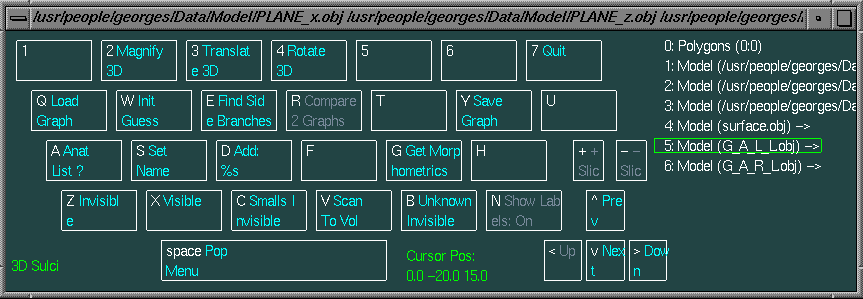
The available labels are listed in the following table (1).
| number | sulcus name | color/scanning_color | scanning_label |
| 0 | unknown_name | BLACK/CYAN | 4 |
| 1 | L_F_Ascending_Ramus | GREEN | 18 |
| 2 | L_F_Horizontal_Ramus | YELLOW | 6 |
| 3 | L_F_Incisura | DARK_BROWN 5 | 14 |
| 4 | L_F_Inferior_Frontal | PURPLE | 7 |
| 5 | L_F_Middle_Frontal | RED | 1 |
| 6 | L_F_Precentral | YELLOW | 6 |
| 7 | L_F_Superior_Frontal | ORANGE | 99 |
| 8 | L_Fo_Intermedius | GREEN | 18 |
| 9 | L_Fo_Lateralis | BLUE | 3 |
| 10 | L_Fo_Medialis | ORANGE | 99 |
| 11 | L_Fo_Olfactory | DARK_BROWN 6 | 14 |
| 12 | L_Fo_Transversus | RED | 1 |
| 13 | L_P_Intraparietal | GREEN | 18 |
| 14 | L_P_Postcentral | BLUE | 3 |
| 15 | L___Central | MAGENTA | 8 |
| 16 | L___Sylvian | RED | 1 |
| 17 | L_T_Superior_Temporal | GREEN | 18 |
| 18 | L_T_Inferior_Temporal | MAGENTA | 8 |
| 19 | L_T_Inferior_Temporal_Ascending_Limb | YELLOW | 6 |
| 20 | L_O_Lateral | RED | 1 |
| 50 | L_F_Subcentral_anterior | DARK_BROWN | 14 |
| 51 | L_F_Frontal_vertical | MAGENTA | 8 |
| 52 | L_F_Precentral_rostral | BLUE | 3 |
| 60 | L_P_Subcentral_posterior | ORANGE | 99 |
| 70 | L_T_Anterior_Collateral | GREEN | 18 |
| 71 | L_T_Posterior_Collateral | MAGENTA | 8 |
| 72 | L_T_Posterior_Ascending_Superior_Temporal | ORANGE | 99 |
| 73 | L_T_Posterior_Horizontal_Superior_Temporal | YELLOW | 6 |
| 99 | L___ERROR | CYAN/BLACK | 40 |
| 100 | L___Side_Branche | WHITE | 41 |
There are basically two situations : i) you want to sart the labelling of a sulgraph file; ii) you want to continue or update the labelling of a sulgraph file.
You can do the following:
Run :
disp.geo final_t1.mnc surface.obj G_A_L_L.obj G_A_R_L.obj
For the left hemisphere, select G_A_L_L.obj in the object list and apply :
Init Guess.
`Enter proba_threshold and depth_threshold:'
enter 0.1 0
(for right hemisphere do the same thing on object number 6, i.e G_A_R_L.obj).
Now you can see that a subset of lines have been colored (other lines are black). All colored lines are the lines that have been labeled based on spatial priors. To figure out was is the label of a curve select this curve with the mouse, use Get Morphometrics and check in the terminal window its name (ex. name : L___Central). Refer to table 1 for correspondance between number/name and colour.
Now you can start to update the actual labeling :
In this case you have already a sulgraph file containing a labelling
and you want to a) see what has been already done; b) continue
the labelling.
Apply Init Guess to the appropriate object (i.e. G_A_L_L.obj, G_A_R_L.obj,
G_A_L_Q.obj or G_A_R_Q.obj).
`Enter proba_threshold and depth_threshold:'
enter: 0.9 0
Here the probability threshold is equal to 0.9. Therefore only curves having a probability vector such that one component is superior to 0.9 are labelled. Considering the fact that Set Name and Add buttons set to 1.0 the associated name's probability , these labels are those which were entered manually (i.e. they correspond to a manual labelling). Then after application of Init Guess with this 0.9 probability threshold, you can see what has been already done. You can then follow the scheme described in the previous paragraph to continue and update this labelling.
Once the labelling has been performed on several sulgraph files, it's possible to compute new SP_AMs using :
~georges/public_bin/mm_sulspam
mm_sulspam is used like this 7
:
mm_sulspam [<options>] <file:list_of_sulgraph> \
<inputfile:mni_template><output_basename:SPAM>
As an example 8:
mm_sulspam proto_list_51.txt \ /data/icbm/mri1/models/average_305_mask_1.00.mnc \ /data/icbm/sbd/seal/Spam/Spams_marita_51/L___Central.mnc \ -clobber -sulind 15 -proba 0.9;
was used to generate the SP_AM of the left central sulcus (-sulind 15) which were manually labelled (-proba 0.9) from 51 manually labeled graphs. ``proto_list_51.txt'' is a text file (the actual file is stored under ~georges/Source/Minctools/Mm_sulspam) which contains the list of input sulgraph files:
proto_list_51.txt /data/icbm/sbd/seal/00100/Sul_graph/lab_marita.sulgraph /data/icbm/sbd/seal/00101/Sul_graph/lab_marita.sulgraph .... /data/icbm/sbd/seal/00157/Sul_graph/lab_marita.sulgraph /data/icbm/sbd/seal/00158/Sul_graph/lab_marita.sulgraph
Figure 4 shows an example of such a SP_AM.
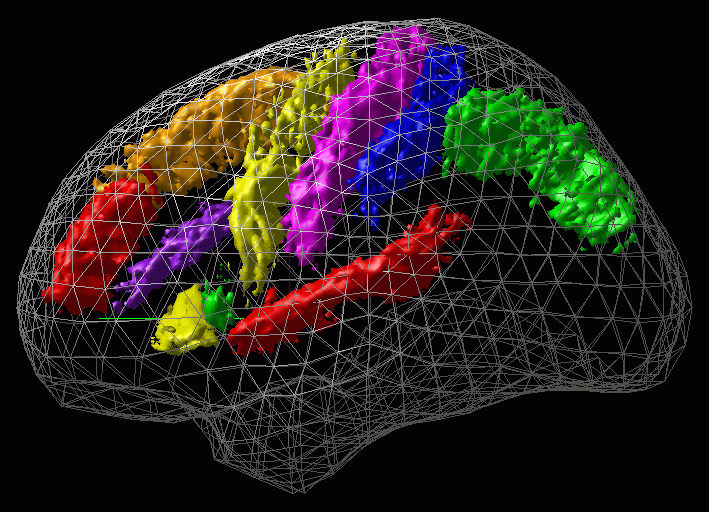
The first phase of the ICBM project had consisted in studying spatial variability of anatomical structure in the MNI-Talairach referential frame in order to construct Statistical Probability Anatomical Maps (SP_AMs) of main anatomical structures . This section present the work done within this context giving also practical informations in order to internally retrieve data (the utimate goal of ICBM being to share these data). A complete list of most recent SP_AMs is stored in : ~georges/Source/Minctools/Include/new_tab_of_SPM.text.
These data were obtained before SEAL was developped. These data represent spatial variability of sulcal superficial traces while SEAL's data (refer to 6.3) represent spatial variability of sulcal median surfaces. I strongly suggest to not use these data as new data are now available (refer to 6.3).
A trained neuroanatomist, blinded as to hemispheric location of the
sulcus by reflection of hemisphere, identified and manually labelled the
superficial trace of a restricted subset of 16 sulci on 51 brains :
(a) the Central Sulcus and (b) the Sylvian Fissure; (c) the Superior, (d)
the Middle, and (e) the Inferior Frontal Sulci; (f) the Ascending and (g)
the Horizontal Rami of the Sylvian Fissure, as well as (h) the Incisura
(located between these other two rami); (i) the Olfactory Sulcus as well
as (j) the Sulcus Lateralis, (k) the Sulcus Medialis, (l) the Sulcus Intermedius,
and (m) the Sulcus Transversus (all found along the orbitofrontal surface
of the brain); (n) the Precentral and (o) the Postcentral Sulci; and finally
(p) the Intraparietal Sulcus. These data were used to construct associated
sulcal SP_AMs, representing the likelihood of finding a particular structure
at each 3-D stereotaxique coordinates. These SPAMs are stored under :
sulcus/Tracing/Sulci/Sulcal_Probability_Maps/
You can find the list of these SP_AMs in the file :
~georges/Source/Minctools/Include/tab_of_SPM.text.
The list of subjects that were labeled can be found in table 2
9.
| a | 00100 | 00101 | 00102 | 00103 | 00104 | 00105 | 00106 | 00107 | 00108 | 00109 |
| b | 00110 | 00111 | 00112 | 00113 | 00114 | 00115 | 00116 | 00117 | 00118 | 00119 |
| c | 00120 | 00121 | 00122 | XXX | 00124 | 00125 | 00126 | 00127 | 00128 | XXX |
| d | XXX | 00131 | XXX | 00133 | 00134 | 00135 | XXX | 00137 | 00138 | 00139 |
| e | 00140 | 00141 | 00142 | 00143 | 00144 | 00145 | 00146 | XXX | 00148 | 00149 |
| f | 00150 | 00151 | 00152 | XXX | XXX | 00155 | 00156 | 00157 | 00158 | XXX |
We have also at our disposal several SP_AMs that were computed from a set of sulgraph files labeled by Emmanuel PROCYK (in collaboration with Michael PETRIDES). These SP_AMs correspond to the frontal lobe sulci : middle frontal s., superior frontal s., inferior frontal s. and also the precentral s., the medial precentral s., the central s., the subcentral anterior sulcus, the subcentral posterior sulcus. 51 brains were studied and the associated sulgraph were labeled. The file /data/icbm/sbd/seal/README contains the list of subjects that were labeled (it is the same list as the one given in table 2). Each labeling is stored in a /data/icbm/sbd/seal/00xxx/Sul_graph/lab_manu_auto.sulgraph.gz file. Computed SP_AMs are no more available because this labelling was updated by Marita SCHMITT.
The labels of Caramanos and Emmanuel Procyk were then transfered to
the lab_marita.sulgraph files and checked by Marita.
This set of 51 sulgraph files contains the most complete sulgraph
labeling. These files are under
/data/icbm/sbd/seal/00XXX/Sul_graph/lab_marita.sulgraph . Associated SP_AMs
are stored under /data/icbm/sbd/seal/Spam/Spams_marita_51. These SP_AMs
represent the data that should be available within the ICBM project. A
complete list of most recent SP_AMs is stored in :
~georges/Source/Minctools/Include/new_tab_of_SPM.text.
These SP_AMs represent the 3D distribution of the sulcus median surface (not just the superficial trace distribution as done by Caramanos).
The file ~georges/Source/Minctools/Include/new_tab_of_SPM.tex
contains the filename and path of the SP_AMs.
Most of the SP_AMs are under : /data/icbm/sbd/seal/Spam/Spams_marita_51
Some of them are also under :
/data/npsych/home/sulcus/Tracing/Sulci/Sulcal_Probability_Maps
Unfortunatly this is not straightforward as there are several modifications to make in the code. Therefore the official ``maintainer'' will have to update the code and recompile the library LibSul, Display.geo and then mm_suldata.
case R\_\_\_NotSetYet\_1 : color = YOURCOLOR;
break;
In the case your want to use a new colour (which was not previously used, i.e. which does not appear in table 1), in:
int compute_label_from_color(Colour col, Colour
*scanning_col)
put this lines :
if(col == YOURCOLOR){
label = YOURLABEL;
*scanning_col = YOURSCANNINGCOLOR;
found = YES; //DIFFER
}
Note that the three last actions have no effect on the code, but you need to make them in order to keep everything updated.
Once this is done, recompile the library LibSul, the program mm_suldata (make install) and Display.geo. To recompile Display.geo, you need to do the following (in order to have: setenv SRC_DIRECTORY /data/nil/david/Release) :
cd ~georges/Source/Display source ~david/Source/Makefile_multi_architecture/cshrc.csh; source ~david/Release/cshrc.csh; make Display.geo;
Again it's the ``maintainer'' who should do this. If a new SP_AM becomes
available you may want to incorporate this information in the computation
of new probability vector (i.e. in the computation of new sulgraph
file). To do that you have to update ~georges/Source/Minctools/Include/new_tab_of_SPM.text.
Example : if the SP_AM of L_T_Superior_Temporal (17) becomes available
edit line 17 of ~georges/Source/Minctools/Include/new_tab_of_SPM.text
(i.e. add the filename there). Edit also line 117 with the corresponding
path for R_T_Superior_Temporal (117). Recompile and install mm_suldata
(i.e. cd ~georges/Source/Minctools/Mm_suldata; make install) .
The new version of sulcus_graph.pl will incorporate these new SP_AMs in
the computation of the probability vectors.
As said before, automatic labelling of the main cortical sulci is done using prior knowledge for the expected 3D spatial distribution of these entities. Probabilistic identification of a new graph is achieved by integrating each sulcal spatial distribution map over a sulcus' median surface and assigning it the sulcus name corresponding to the maximum integrated probability value. Quantitative evaluation of the automatic labelling accuracy with regards to the different types of errors (under-estimation of sulcal extend, confusion between adjacent sulci) were computed. Results show that correct identification of eight main cortical sulci studied can be reached in 34 % to 70 % of the cases depending on the level of accuracy needed. For more information on this problem, refer to ~georges/Docs/Postdoc/Article/article.tex and ~georges/Docs/Postdoc/Article/Script.
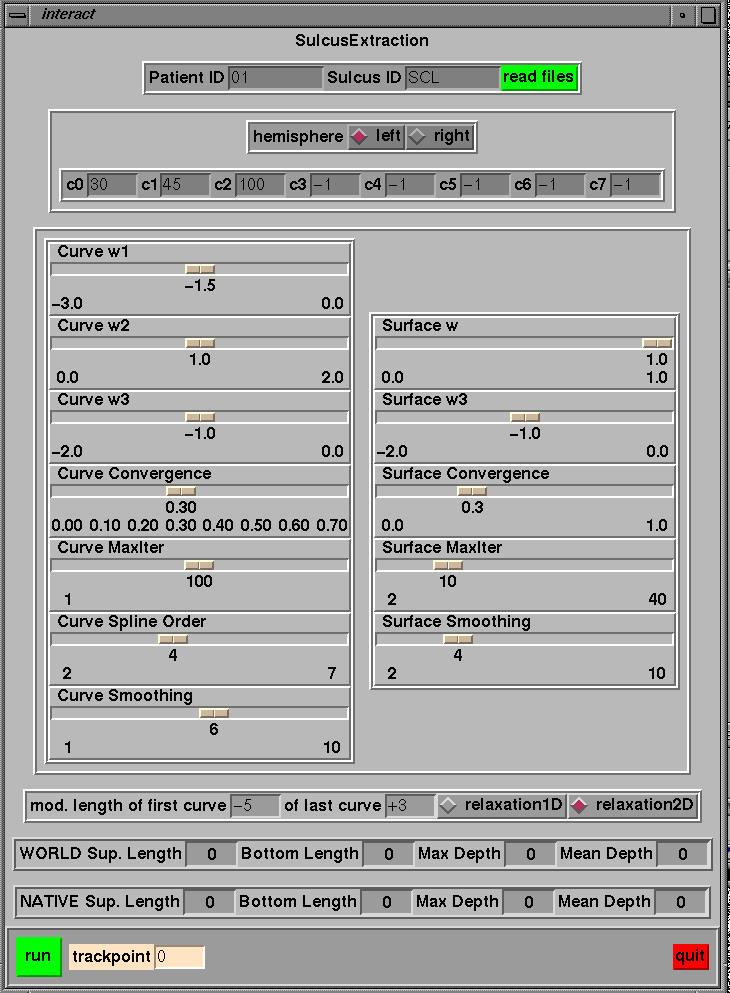
Post-segmentation can be done using ~georges/Source/Minctools/Mm_depth_interac/Tcl/interact. The reason for doing a post-segmentation is that the structural decomposition of the sulcal topography as obtained by SEAL is based on the observation of the external part of the brain. Then a sulcus is generally decomposed in several segments (several portions). Then if it's needed to model a sulcus as a single element, one need to link the different portions of the sulcus of interest. The following illustration tells the post-segmentation to connect curve number 30, 45 and 100 (of the left hemisphere) to create a simple surface. Moreover the user wants to cut 5 mm at the beginning of the curve and want to add 3 mm at the end of the fold. The user can also change the parameters of the active ribbon. Note that this program need to know where data are stores (i.e. where to find POT1, POT2 etc...), this is defined in interact.h.
http://www.bic.mni.mcgill.ca/~georges/publications/publi.html
~georges/Docs/Postdoc/TMI/Submission/le_goualher_149_98_IEEE_TMI.ps.gz
~georges/Docs/Postdoc/Article/article.tex
~georges/Docs/Postdoc/Argenti/Paper1/Paper_two/Submission/goualher_neuroimage_main.ps.gz
~georges/Data_nil/Docs/Postdoc/Repport1/rep_oct98.ps.gz
/data/icbm/sbd/seal/Slides
summary: /data/icbm/sbd/seal/Slides/MNI/summary.sho
figures: /data/icbm/sbd/seal/Image_Rapport